3D Brain Segmentation
Subcortical structure segmentation of 3D brain MRIs
This research was done as a part of the machine learning research scientist internship at Owkin. It focuses on probing the limits of the nnUNet pipeline in low-data regimes and comparing it to SynthSeg, a synthetic-data-based segmentation approach.
Results
Experiments were performed using nnUNet and SynthSeg. SynthSeg is treated as an out-of-the-box benchmark requiring no training, while nnUNet serves as a data-driven baseline trained with very limited annotations (5 or 30 image–mask pairs).
Quantitative results are summarized below, followed by representative visualizations. For full experimental details and extended analysis, the reader is referred to the complete report.
Performance comparison
| Model | OASIS DSC | OASIS HD95 | OASIS NSD | ADNI DSC | ADNI HD95 | ADNI NSD |
|---|---|---|---|---|---|---|
| nnUNet5 | 82.53 | 5.52 | 1.97 | 68.54 | 14.43* | 7.75* |
| nnUNet30 | 88.97 | 1.58 | 29.15 | 84.85 | 4.72 | 2.05 |
| SynthSeg | 76.59 | 3.69 | 0.93 | 69.13 | 2.94 | 1.24 |
nnUNet trained on 5 images
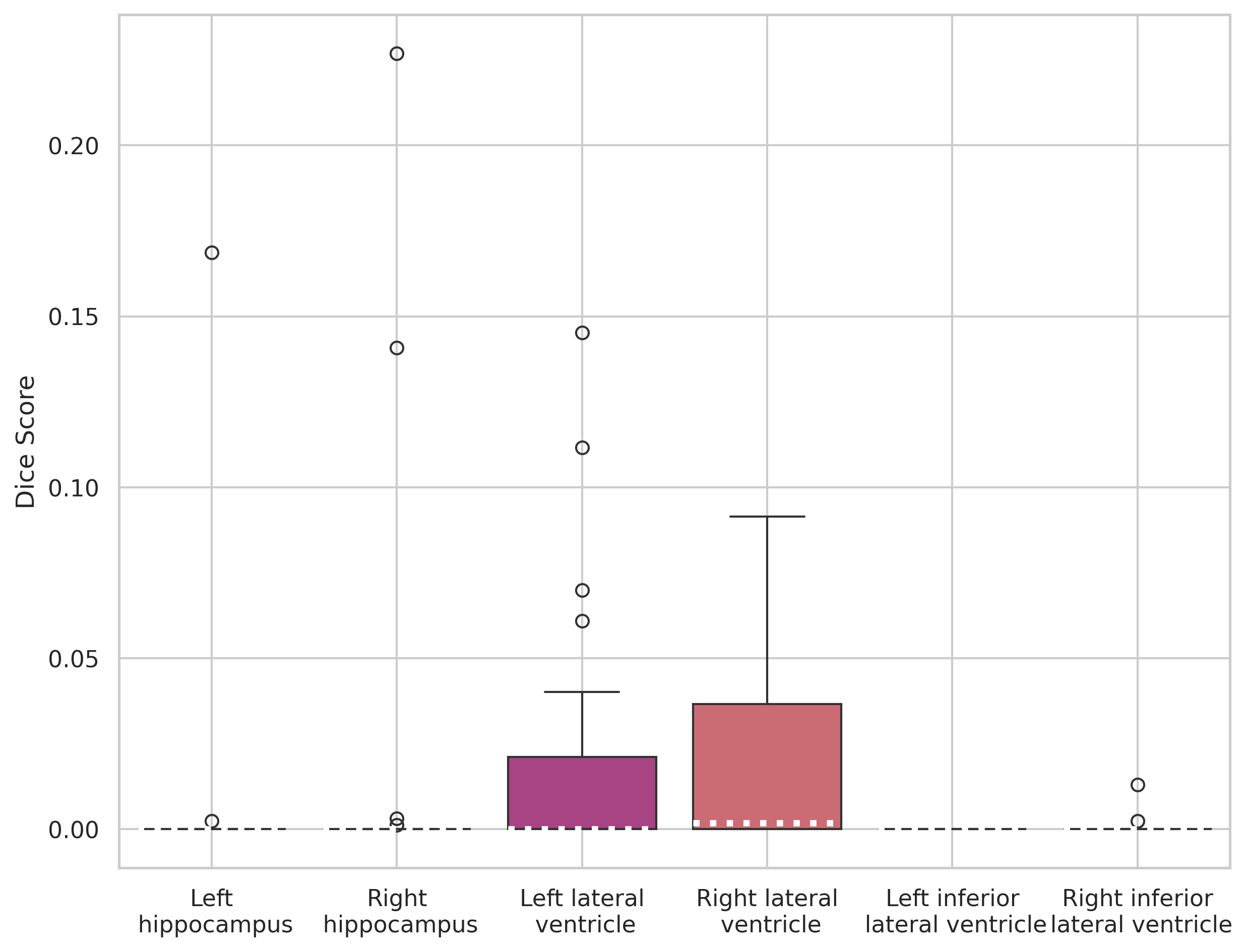
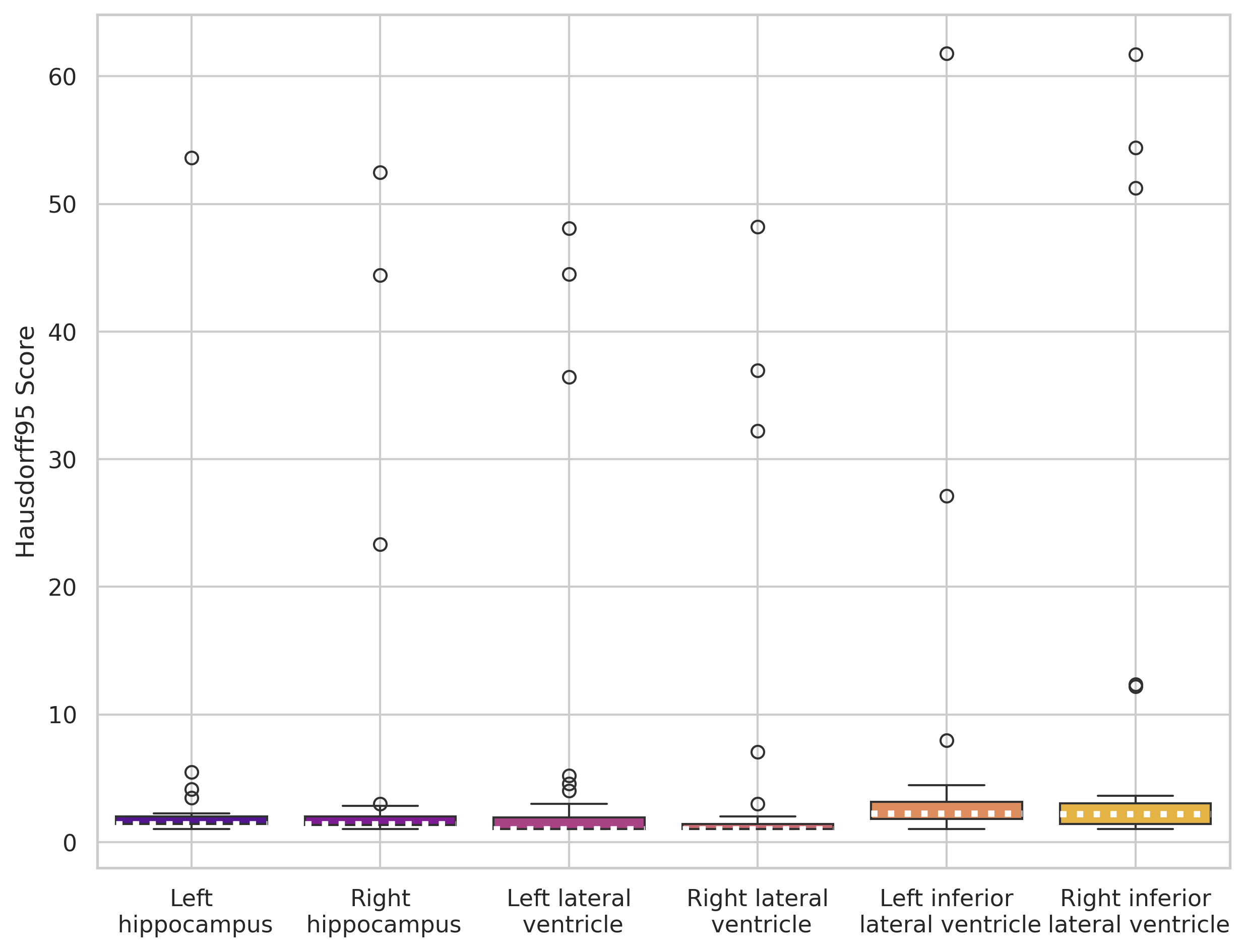
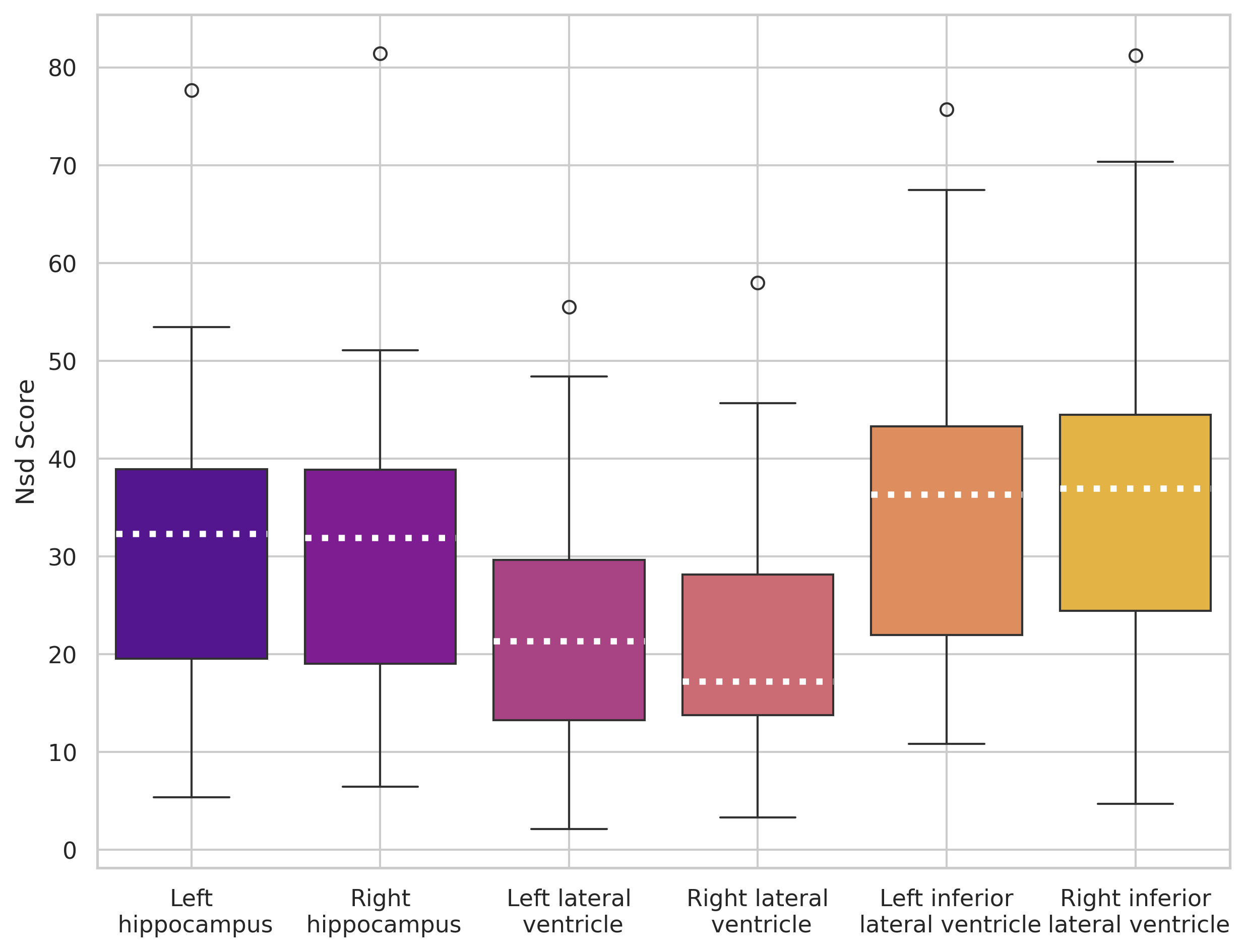
nnUNet trained on 30 images
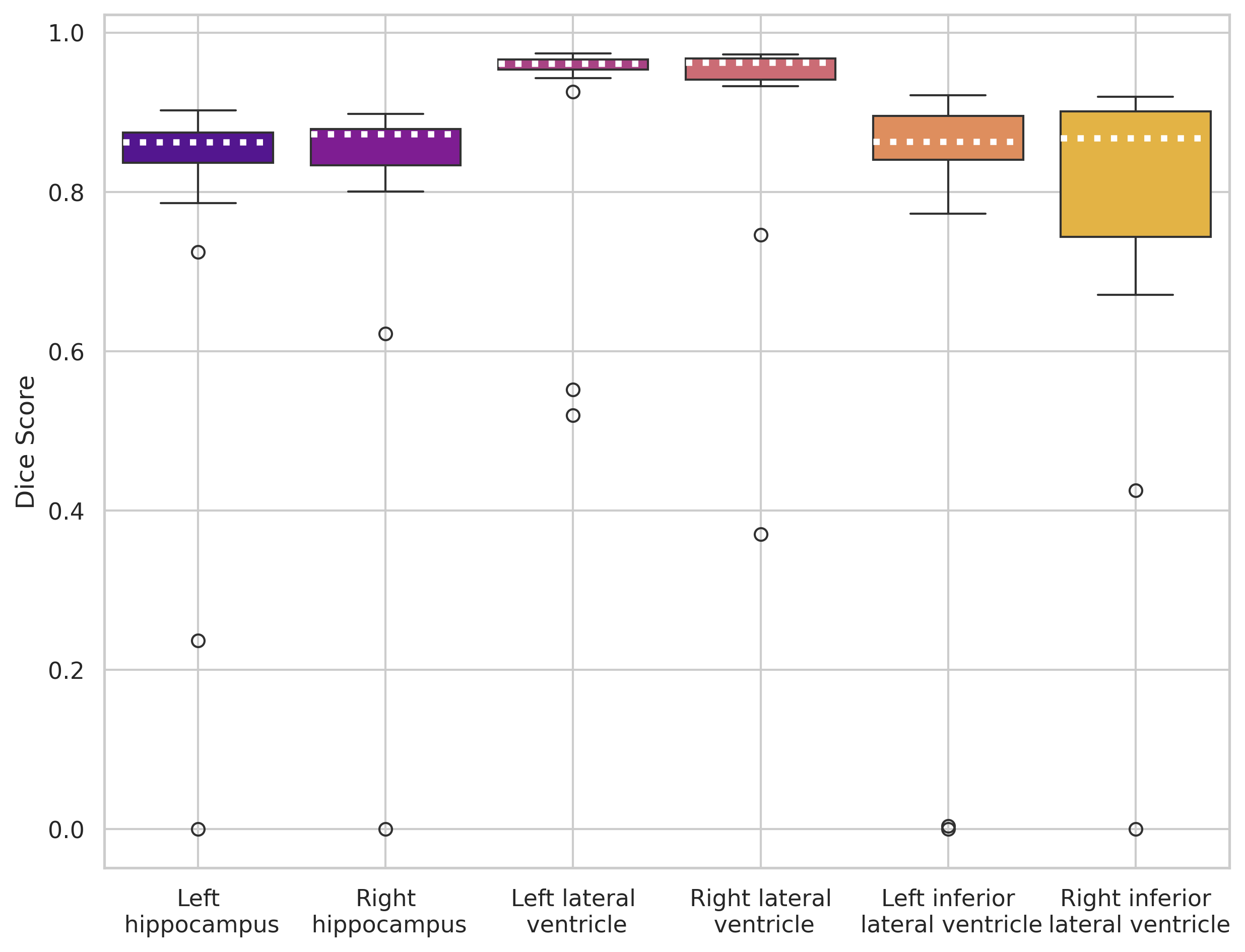
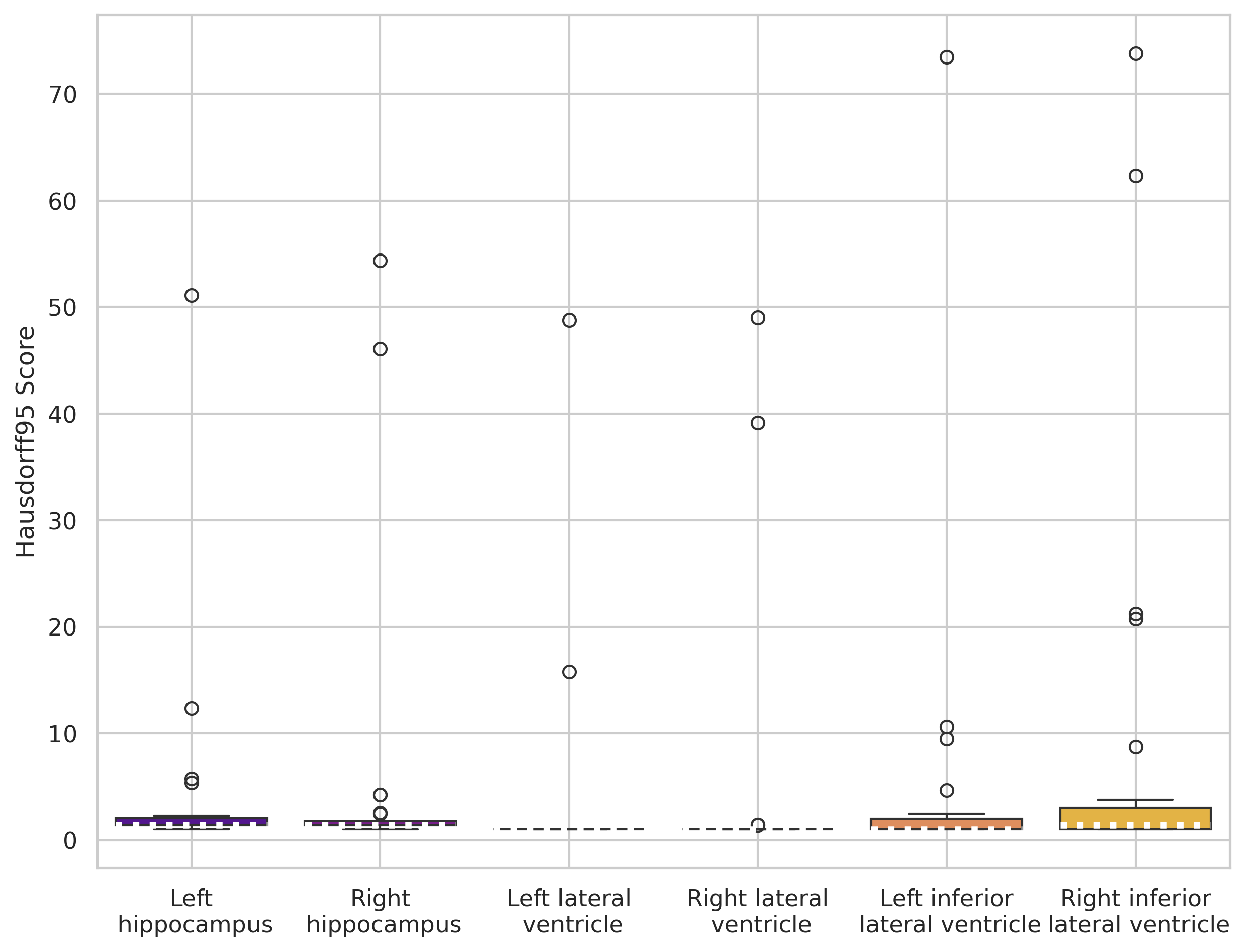
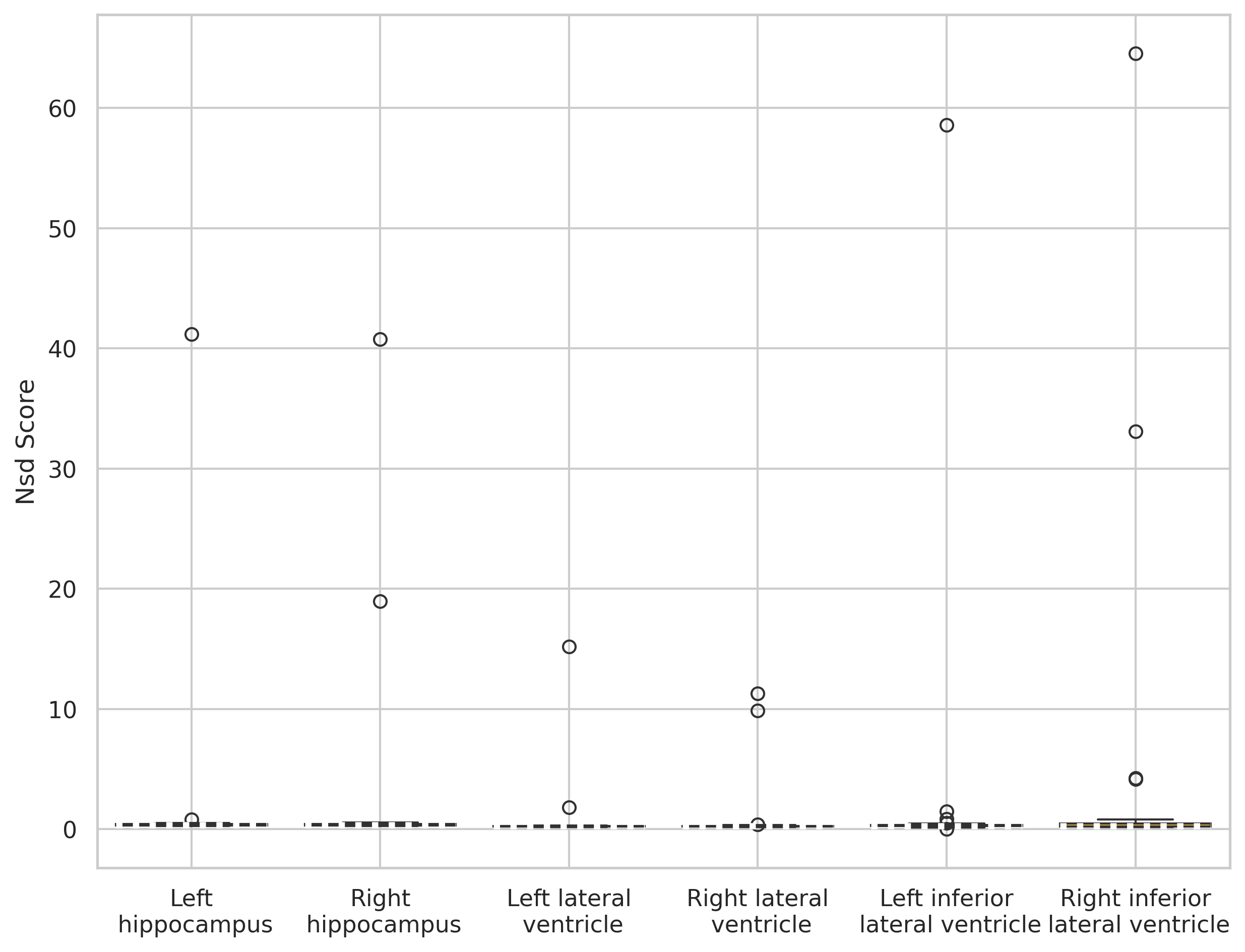
SynthSeg (out-of-the-box)
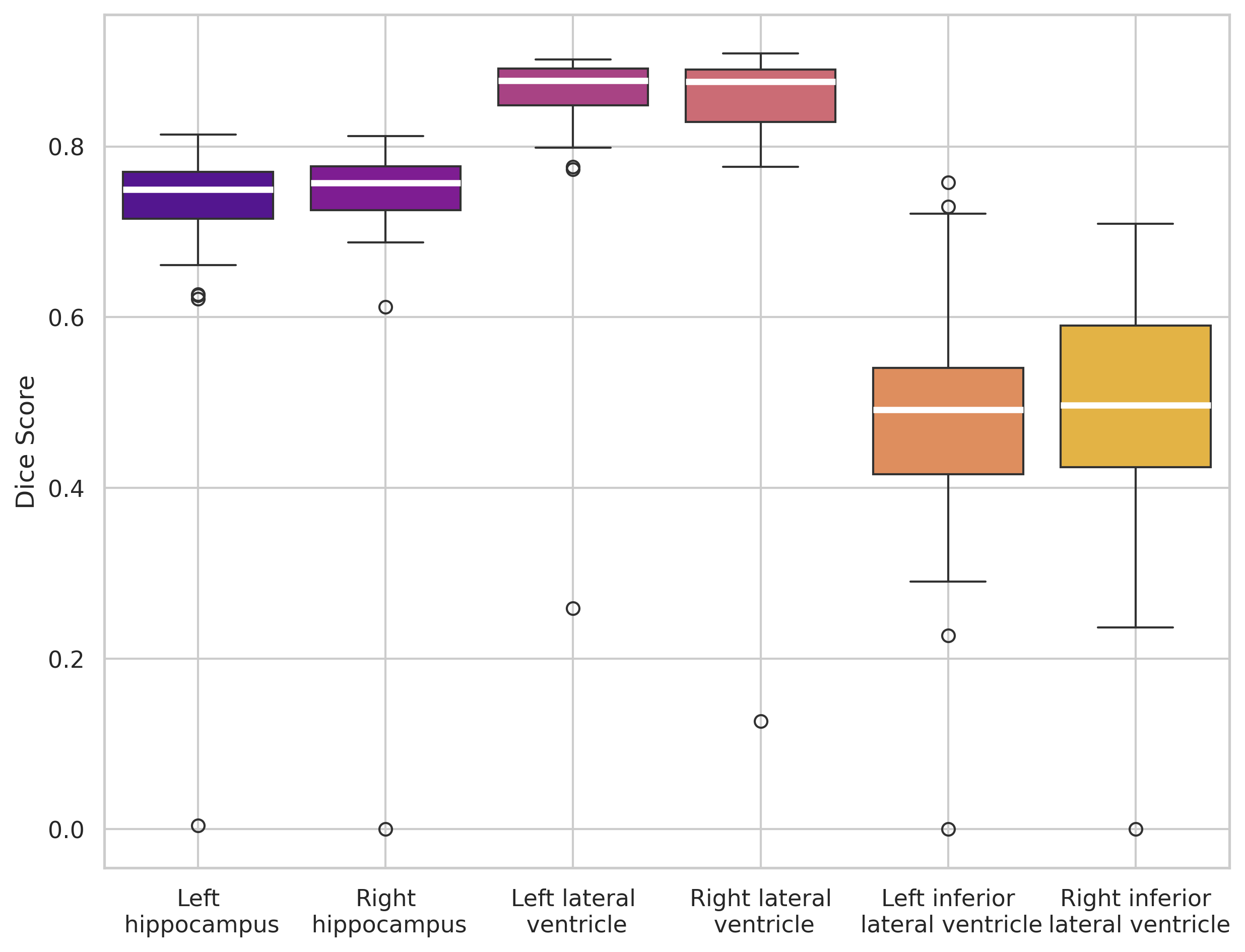
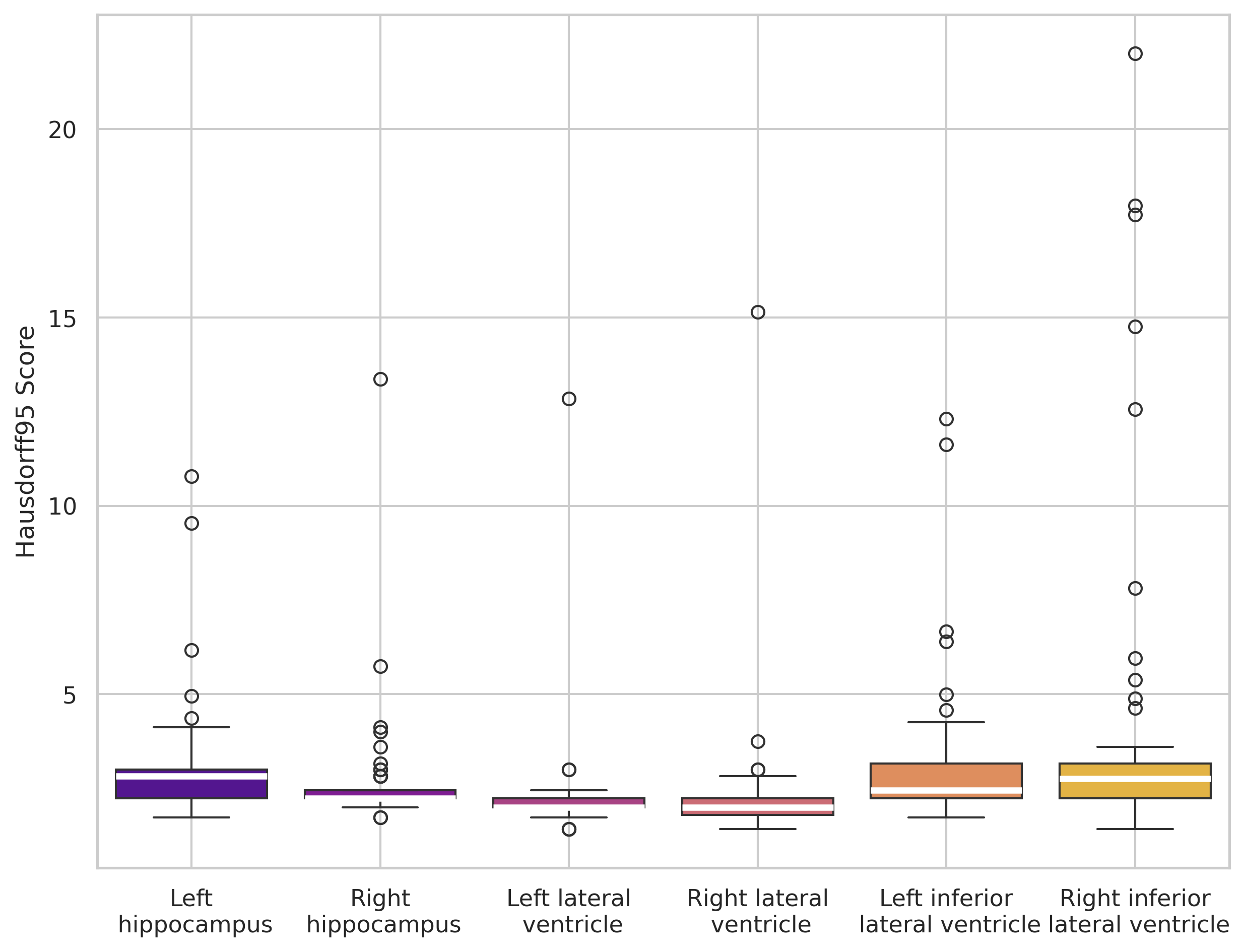
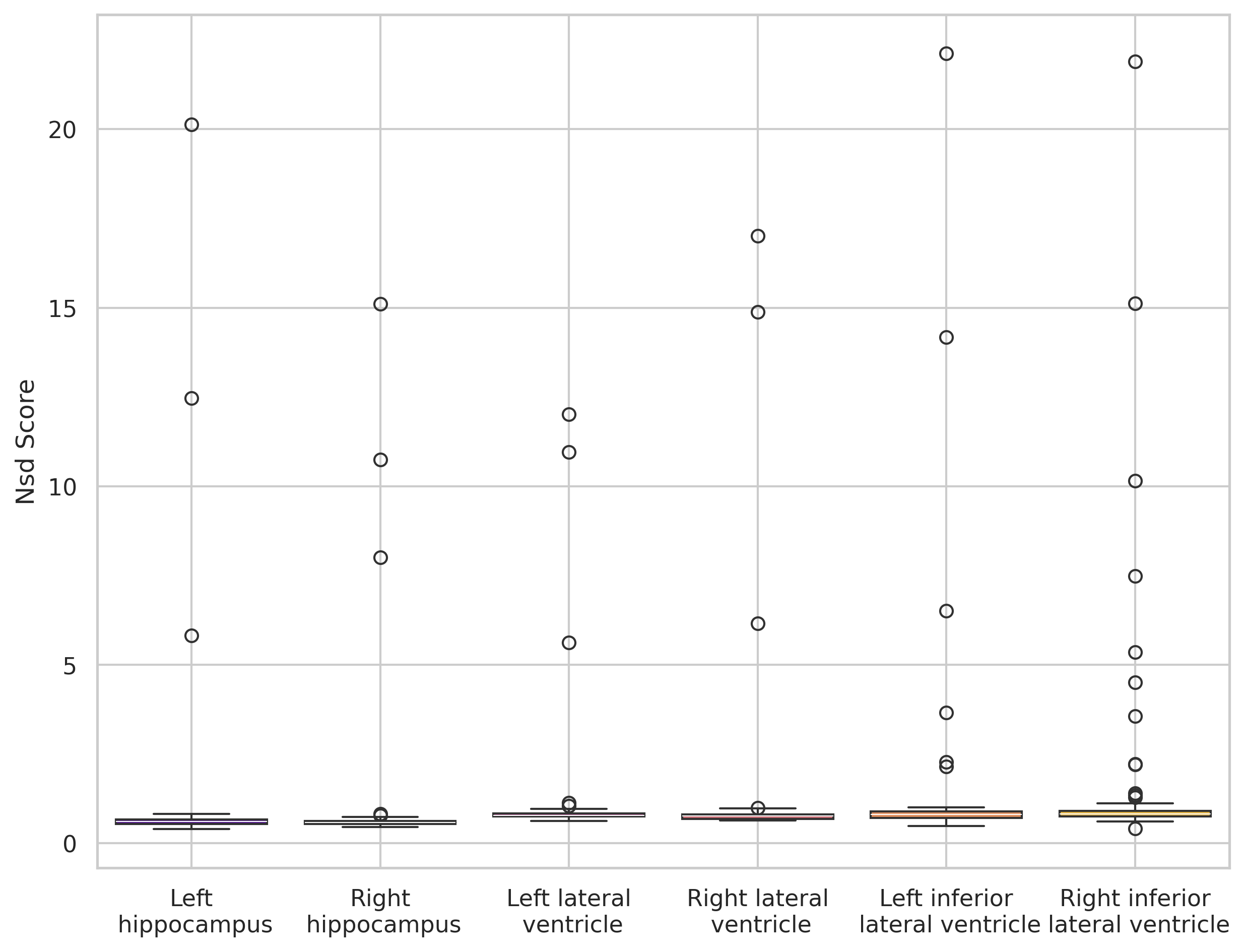
Symmetry effect and qualitative analysis
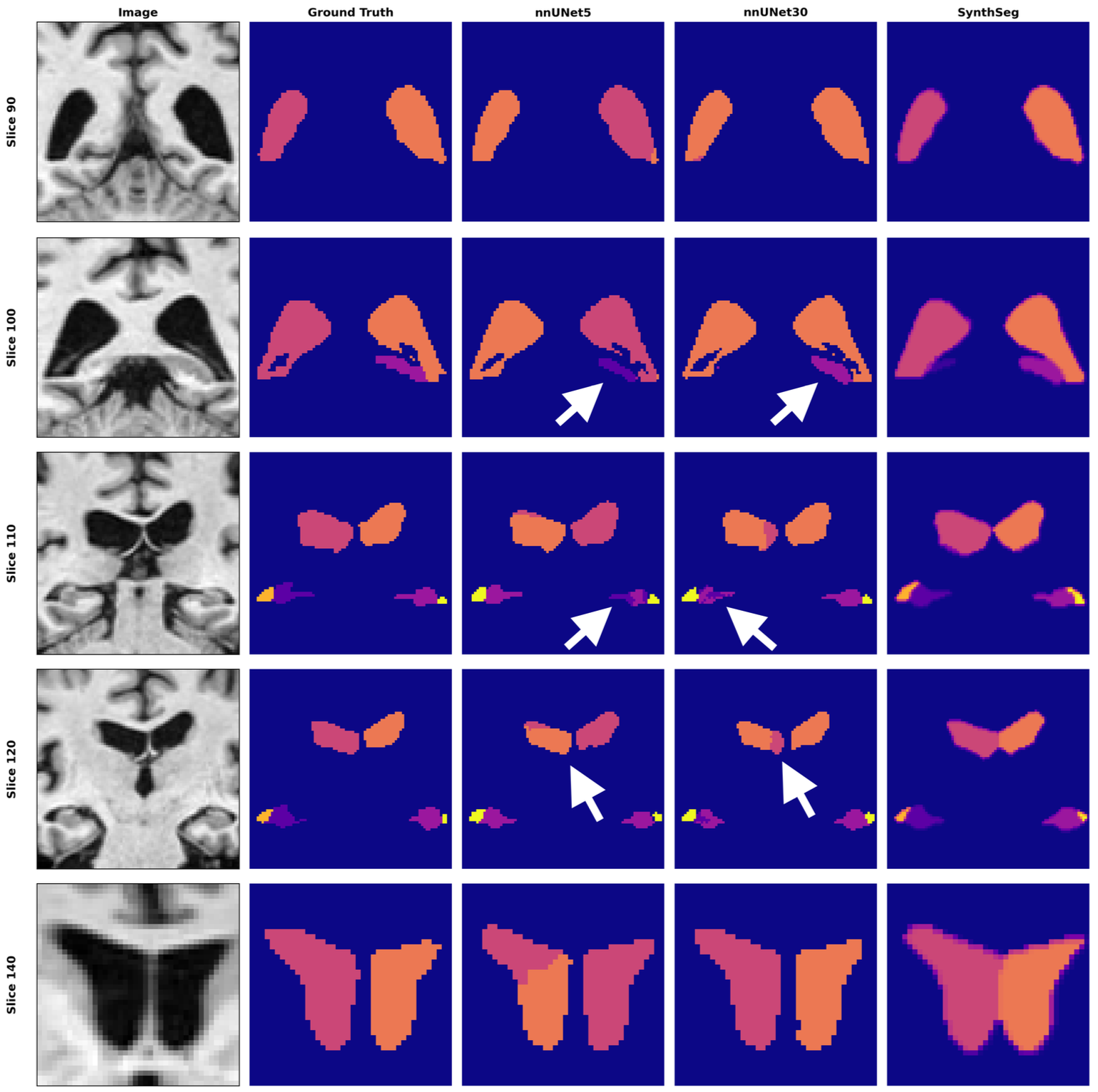
nnUNet cross-dataset inference reveals a strong bias toward symmetric structure placement: shapes are captured correctly, but left/right class assignment fails. Dice scores increase dramatically when labels are merged by structure or foreground, indicating preserved shape understanding but poor semantic consistency across domains.
This page presents a curated visual summary. For full metrics, ablations, and discussion, please refer to the complete technical report.